I have spent the last decade working as a data developer, and if there is one tool I use every single day, it is the Pandas library.
When I first started analyzing US economic data, I realized that getting the data into Python was often the hardest part of the job.
Most of the time, your data arrives in a Comma-Separated Values (CSV) format, which seems simple until you encounter messy formatting.
In this guide, I will share my firsthand experience using the read_csv function to handle everything from small retail files to massive census datasets.
How to Use the Basic read_csv Function
The most common task you will face is simply loading a local file into a DataFrame so you can start your analysis.
In my experience, I usually keep my Python script in the same folder as my data to keep things organized.
Suppose you have a file named us_retail_sales.csv that contains monthly revenue figures for a chain of stores across the United States.
Here is the code I use to bring that data into my environment:
import pandas as pd
# I am loading a standard US retail sales dataset
df = pd.read_csv('us_retail_sales.csv')
# I always check the first few rows to ensure it loaded correctly
print(df.head())You can refer to the screenshot below to see the output.
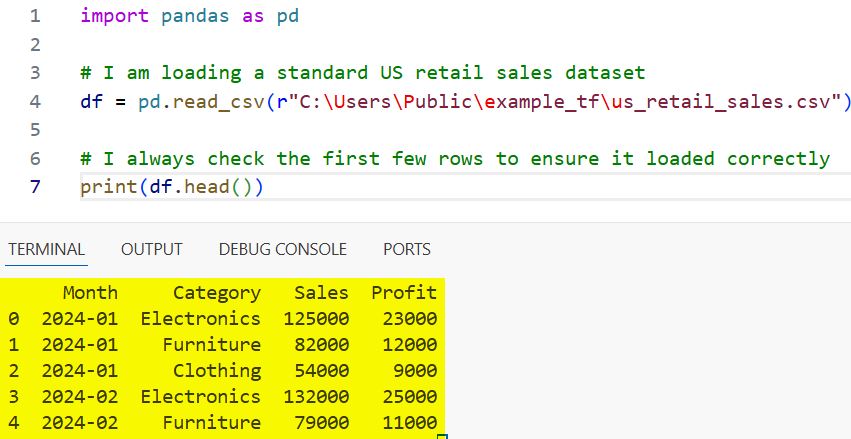
When you run this, Pandas automatically identifies the header row and assigns the data types for each column.
I find this “auto-detection” works about 90% of the time, but the other 10% requires a bit more professional finesse.
Read CSV Files with Custom Delimiters
Sometimes, you might receive a file that uses something other than a comma to separate the data points.
I often see this with older US government export files, where tabs or semicolons are used to prevent errors with address fields.
If you try to read a tab-separated file with the default settings, Pandas will put all the data into a single, messy column.
I learned early on that you must explicitly tell Pandas what the separator is by using the sep parameter.
import pandas as pd
# In this example, I am reading a file where data is separated by tabs
# This is common in US real estate listing exports
df = pd.read_csv('us_listings.txt', sep='\t')
print(df.head())You can refer to the screenshot below to see the output.
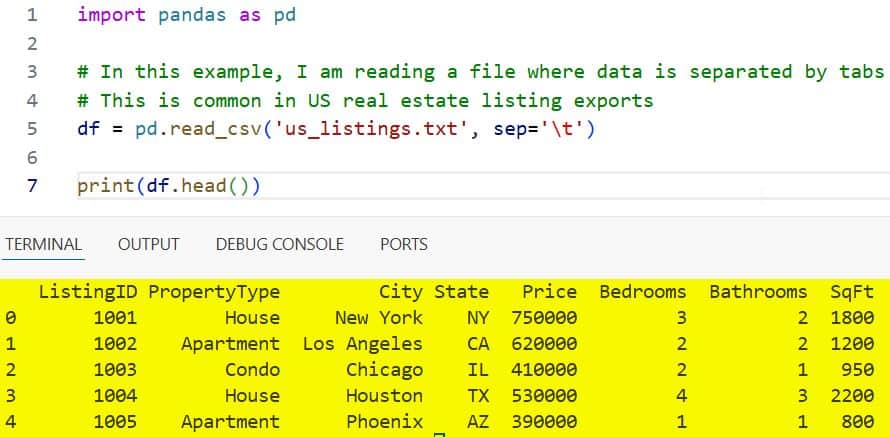
By using sep=’\t’, I am ensuring that the residential addresses and price points are split into the correct columns.
It is a small tweak, but it saves me hours of manual data cleaning later in the workflow.
Select Specific Columns to Save Memory
When I work with large US Census datasets, I rarely need every single one of the hundreds of columns provided.
Loading unnecessary data into your computer’s memory is a quick way to crash your Python script.
I prefer to use the usecols argument to grab only the specific data points that matter for my current project.
For instance, if I only care about the state name and the total population, I tell Pandas to ignore everything else.
import pandas as pd
# I am only interested in the 'State' and 'Population_2024' columns
# This saves a significant amount of RAM when dealing with large files
selected_columns = ['State', 'Population_2024']
df = pd.read_csv('us_census_data.csv', usecols=selected_columns)
print(df.info())You can refer to the screenshot below to see the output.
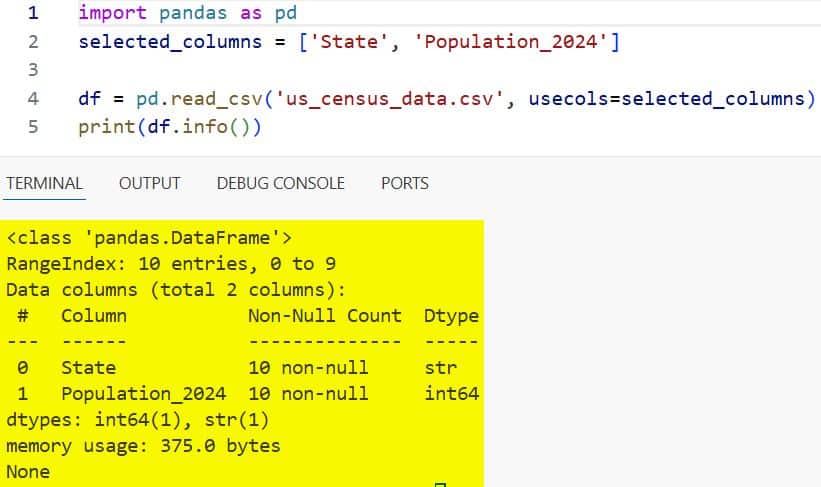
This approach makes my code run much faster, and it keeps my workspace clean and focused on the task at hand.
I highly recommend this habit if you are working on a machine with limited memory or dealing with “Big Data.”
Handle Headers and Index Columns
Not every CSV file comes with a perfect header row at the very top of the document. I have often received financial reports from US banks where the first few lines are just “Fine Print” or legal disclaimers.
If you don’t account for this, Pandas will try to use that legal text as your column names, which creates a total mess.
I use the header parameter to tell the script exactly which row contains the actual column titles.
import pandas as pd
# Here, I am telling Pandas that the actual data starts on the third row (index 2)
# I am also setting the 'Transaction_ID' as my primary index
df = pd.read_csv('bank_statement.csv', header=2, index_col='Transaction_ID')
print(df.head())You can refer to the screenshot below to see the output.
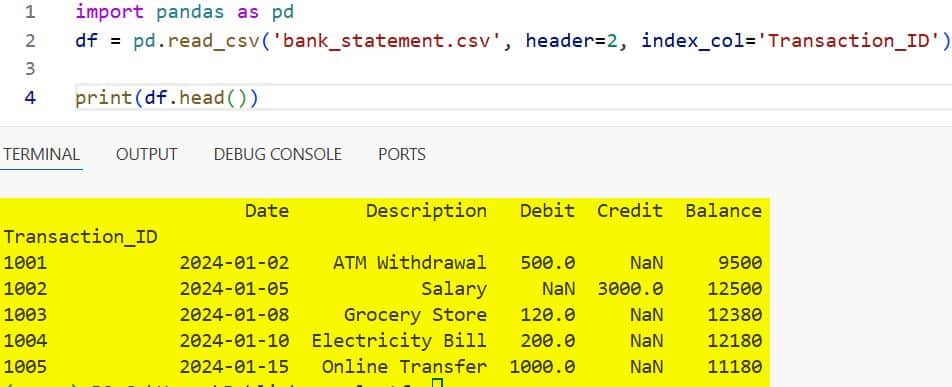
Setting an index column like Transaction_ID makes it much easier to look up specific records later on.
It transforms the DataFrame from a simple list into a powerful, searchable database table.
Correctly Parsing Dates during Import
One of the biggest headaches I face as a developer is dealing with date formats in US-based datasets.
By default, Pandas sees dates like “05/12/2024” as simple strings of text rather than actual time objects.
This means you cannot easily perform time-series analysis or filter by month without converting them first.
I save myself a step by using the parse_dates parameter directly inside the read_csv function.
import pandas as pd
# I am importing US stock market data and converting the 'Date' column immediately
df = pd.read_csv('nasdaq_prices.csv', parse_dates=['Date'])
# Now I can check the data types to confirm 'Date' is a datetime object
print(df.dtypes)Once the dates are parsed correctly, I can perform operations like calculating the “Year-over-Year” growth in seconds.
It is a simple trick that I wish someone had told me when I first started using Python for finance.
Manage Missing Values and NaNs
In the real world, data is rarely perfect, and you will often find empty cells in your CSV files.
I frequently see this in US healthcare surveys, where certain questions were left unanswered by the participants.
Pandas usually marks these as NaN (Not a Number), but sometimes you need to specify what counts as “missing” data.
If a dataset uses placeholders like “Unknown” or “N/A”, I tell Pandas to treat those as null values immediately.
import pandas as pd
# I am defining a list of values that should be treated as missing
# This is helpful for cleaning up messy US survey data
missing_vals = ['n/a', 'unknown', '--']
df = pd.read_csv('healthcare_survey.csv', na_values=missing_vals)
# I can then count how many missing values exist in each column
print(df.isnull().sum())Identifying these gaps early allows me to decide whether to fill them with averages or drop them entirely.
Clean data leads to accurate insights, and this is the first step in that quality control process.
Skip Rows and Footers
Sometimes a file contains extra information at the bottom, like a summary total or a “Produced by” timestamp.
If Pandas tries to read these summary rows as standard data, it will often cause errors in your numeric columns.
I use the skiprows and skipfooter parameters to trim the “fat” from the top and bottom of my files.
When using skipfooter, I always make sure to set the engine to ‘python’ to avoid any compatibility warnings.
import pandas as pd
# I am skipping the first 5 rows and the last 2 rows of the file
# This is perfect for US government reports that have long intros and outros
df = pd.read_csv('federal_spending.csv', skiprows=5, skipfooter=2, engine='python')
print(df.tail())This ensures that only the raw data rows are included in my analysis, making the results much more reliable.
It is a professional touch that ensures your automated scripts don’t break when the report format changes slightly.
Handle Large Files with Chunking
If you ever try to load a multi-gigabyte file containing US transportation records, your computer might freeze.
In these situations, I never load the whole file at once; instead, I process it in smaller “chunks.”
I use the chunksize parameter, which returns an object that I can loop through one piece at a time.
This allows me to calculate totals or averages without ever overwhelming my system’s memory.
import pandas as pd
# I am reading a massive dataset in blocks of 10,000 rows each
# This is how I handle heavy US logistics and shipping data
chunk_size = 10000
batch_no = 1
for chunk in pd.read_csv('us_freight_movements.csv', chunksize=chunk_size):
print(f'Processing Batch {batch_no}...')
# Here I could perform calculations on each chunk
batch_no += 1This method is the gold standard for high-performance data engineering in Python. It turns a task that would be impossible on a laptop into a routine operation that finishes in minutes.
Read Data Directly from a URL
One of my favorite features is the ability to bypass the “download” button entirely. Many US agencies, like the Bureau of Labor Statistics, provide direct links to their CSV files online.
I can pass that URL directly into read_csv, and Pandas will download and load the data in one go.
This is incredibly useful for creating dashboards that always show the most up-to-date information.
import pandas as pd
# I am fetching the latest US unemployment data directly from an online source
url = 'https://raw.githubusercontent.com/datasets/gdp/master/data/gdp.csv'
df = pd.read_csv(url)
print(df.head())This streamlines my workflow and ensures that I am always working with the freshest data available.
Just make sure you have a stable internet connection before running a script that relies on external URLs.
Specify Data Types for Efficiency
By default, Pandas guesses which data types to use, but sometimes it chooses poorly. For example, it might store a column of US Zip Codes as integers, which removes the leading zeros.
I always define my data types manually using the dtype parameter to prevent this from happening.
Storing data as the correct type also reduces the amount of memory your DataFrame occupies.
import pandas as pd
# I am ensuring Zip Codes are kept as strings to preserve leading zeros
# This is vital for accurate US geographical mapping
data_types = {
'Zip_Code': str,
'Sales_Amount': float
}
df = pd.read_csv('store_locations.csv', dtype=data_types)
print(df.dtypes)
print(df['Zip_Code'].head())This level of control is what separates a beginner from an experienced data professional. It ensures that your data remains accurate and your analysis remains precise from start to finish.
Working with CSVs in Pandas is one of those skills that you will use throughout your entire career.
While the basic command is simple, mastering the parameters I’ve discussed here will make you much more efficient.
I recommend practicing with a few different US-based datasets to see how these settings affect your results.
Once you get the hang of it, you will be able to handle even the messiest files with total confidence.
You may also like to read:
- How to Read Excel Files in Pandas
- How to Compare Two Pandas DataFrames in Python
- How to Use Pandas Concat with Ignore Index
- How to Use Pandas GroupBy Aggregation

Bijay Kumar is an experienced Python and AI professional who enjoys helping developers learn modern technologies through practical tutorials and examples. His expertise includes Python development, Machine Learning, Artificial Intelligence, automation, and data analysis using libraries like Pandas, NumPy, TensorFlow, Matplotlib, SciPy, and Scikit-Learn. At PythonGuides.com, he shares in-depth guides designed for both beginners and experienced developers. More about us.

