One of the most frequent tasks I handle as a Python developer is bringing external data into a Pandas DataFrame for analysis.
While we often talk about CSVs, a lot of legacy data or system logs are stored in plain text files (.txt).
In this tutorial, I will show you exactly how I use Pandas to read text files efficiently, covering different delimiters and common formatting hurdles.
The Go-To Method: Use pd.read_csv for Text Files
Even though the function is named read_csv, it is actually the most versatile tool for reading almost any delimited text file.
I use this method about 90% of the time because it is highly optimized for performance.
Example: Read a Pipe-Delimited Texas Real Estate File
Let’s say you have a text file containing real estate data from Austin, Texas, where the values are separated by a pipe (|) symbol instead of a comma.
import pandas as pd
# I prefer using read_csv even for .txt files by specifying the separator
file_path = 'austin_real_estate.txt'
# Creating a sample text file for the demonstration
with open(file_path, 'w') as f:
f.write("Property_ID|Neighborhood|Price_USD\n")
f.write("101|Zilker|850000\n")
f.write("102|Tarrytown|1200000\n")
f.write("103|Bouldin Creek|975000\n")
# Reading the pipe-delimited text file
df = pd.read_csv(file_path, sep='|')
print(df)I executed the above example code and added the screenshot below.
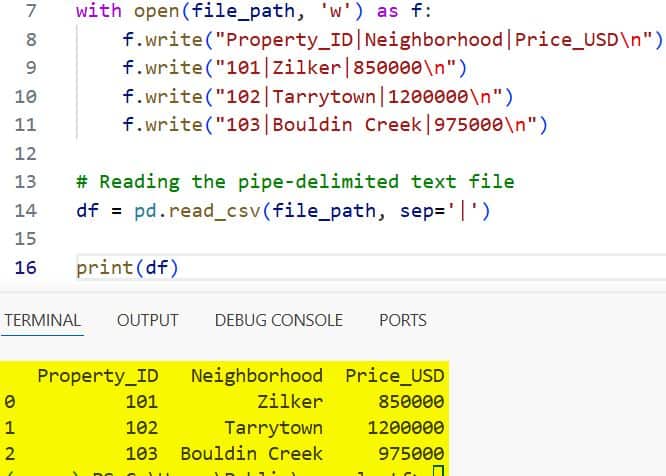
In this code, the sep=’|’ argument tells Pandas exactly how to split the strings into columns. If your file uses tabs, you would simply use sep=’\t’.
Use pd.read_table for Tab-Separated Data
In my early days of coding, I used pd.read_table quite often because its default separator is a tab.
While it functions almost identically to read_csv, it feels more “natural” when dealing with specific .dat or .txt files that are strictly tab-delimited.
Example: US Census Population Data
Imagine we are looking at population stats for major US cities stored in a tab-separated format.
import pandas as pd
# Reading a tab-separated text file
# Note: read_table defaults to sep='\t'
file_path = 'us_city_population.txt'
with open(file_path, 'w') as f:
f.write("City\tState\tPopulation_2023\n")
f.write("New York\tNY\t8335897\n")
f.write("Los Angeles\tCA\t3822238\n")
f.write("Chicago\tIL\t2665039\n")
df_pop = pd.read_table(file_path)
print(df_pop.head())I executed the above example code and added the screenshot below.
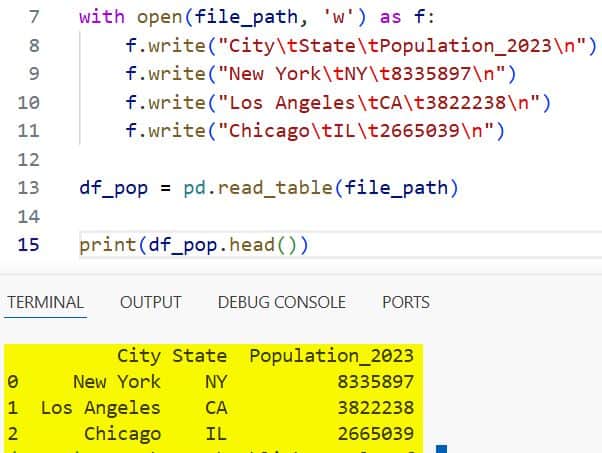
I find this method very clean for scientific datasets or academic exports that frequently use tabs.
Deal with Files That Have No Header
It is quite common to receive system-generated logs that don’t include column names in the first row.
If you don’t tell Pandas this, it will mistakenly treat your first row of data as the header.
Example: Florida Temperature Logs
In this scenario, I’ve got daily high temperatures from Miami, but the file starts right with the data.
import pandas as pd
file_path = 'miami_temps.txt'
with open(file_path, 'w') as f:
f.write("2024-01-01,82\n")
f.write("2024-01-02,84\n")
f.write("2024-01-03,81\n")
# Use header=None and provide names manually
column_names = ['Date', 'Temp_Fahrenheit']
df_weather = pd.read_csv(file_path, header=None, names=column_names)
print(df_weather)I executed the above example code and added the screenshot below.
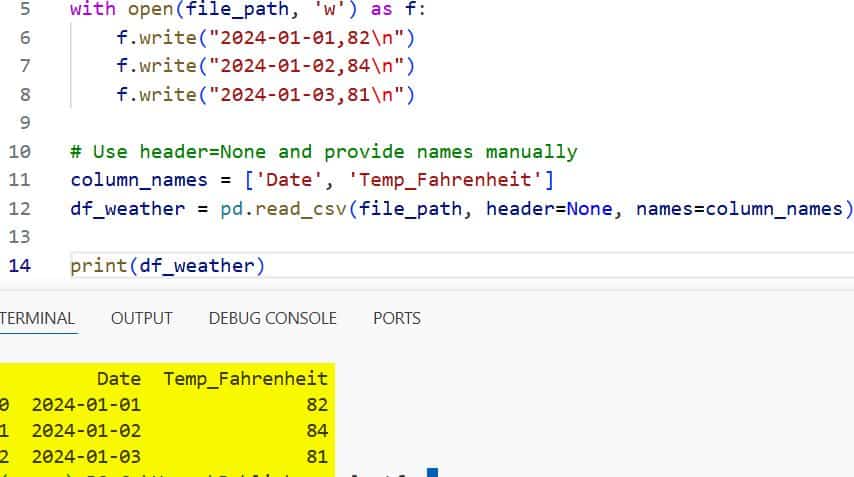
By setting header=None, I ensure every line in the text file is treated as a data record.
Handle Fixed-Width Formatted Text Files
Occasionally, I encounter legacy data from US government agencies or financial institutions that use “Fixed Width” formatting.
In these files, columns aren’t separated by characters like commas; instead, each column occupies a specific number of character spaces.
Example: Read NYSE Stock Data
For this, I use pd.read_fwf, which stands for “Fixed Width File.”
import pandas as pd
file_path = 'stock_ticker_data.txt'
# Each column has a strictly defined width
with open(file_path, 'w') as f:
f.write("AAPL Apple Inc. 185.20\n")
f.write("MSFT Microsoft Corp 405.15\n")
f.write("TSLA Tesla Inc. 175.50\n")
# Pandas is surprisingly good at inferring these widths automatically
df_stocks = pd.read_fwf(file_path, header=None, names=['Ticker', 'Company', 'Price'])
print(df_stocks)If the automatic detection fails, I manually pass a list of widths to the widths parameter.
Skip Rows and Footers in Text Files
Sometimes a text file contains metadata or “junk” lines at the top or bottom that aren’t part of the dataset.
I often see this in exports from accounting software where the first few lines are “Report Title” and “Date Range.”
Example: California Tech Salary Survey
import pandas as pd
file_path = 'salary_survey.txt'
with open(file_path, 'w') as f:
f.write("Annual Salary Survey Report\n")
f.write("Confidential Information\n")
f.write("Job_Title,Location,Salary\n")
f.write("Data Scientist,San Jose,165000\n")
f.write("Software Engineer,Palo Alto,180000\n")
f.write("End of Report\n")
# Skip the first two rows and the last row
df_salary = pd.read_csv(file_path, skiprows=2, skipfooter=1, engine='python')
print(df_salary)Note that when using skipfooter, I always specify engine='python' because the default C engine doesn’t support it.
Read Only Specific Columns (Memory Efficiency)
When I’m working with massive text files—like US flight delay data which can have dozens of columns—I don’t want to load everything into memory.
I only pick the columns I actually need using the usecols parameter.
Example: Flight Data Analysis
import pandas as pd
# Imagine a file with 10 columns, but we only want 'Origin' and 'Delay'
file_path = 'us_flights.txt'
with open(file_path, 'w') as f:
f.write("Date,Origin,Dest,Delay_Min,Carrier,TailNum\n")
f.write("2024-03-01,JFK,LAX,15,Delta,N123\n")
f.write("2024-03-01,ORD,SFO,0,United,N456\n")
df_flights = pd.read_csv(file_path, usecols=['Origin', 'Delay_Min'])
print(df_flights)This simple trick can make your scripts run significantly faster and save a lot of RAM.
Handle Mixed Data Types and Dates
Pandas usually does a great job at guessing data types, but sometimes it needs a nudge, especially with US-style dates (MM-DD-YYYY).
Example: E-commerce Orders
import pandas as pd
file_path = 'orders.txt'
with open(file_path, 'w') as f:
f.write("order_id,order_date,amount\n")
f.write("1001,03-15-2024,250.50\n")
f.write("1002,03-16-2024,99.99\n")
# Use parse_dates to convert the string column to actual datetime objects
df_orders = pd.read_csv(file_path, parse_dates=['order_date'])
print(df_orders.dtypes)
print(df_orders)Converting dates immediately upon reading allows me to perform time-series analysis without extra conversion steps later.
Load Large Text Files in Chunks
If you are trying to read a text file that is several gigabytes in size, your computer might freeze.
In such cases, I process the file in “chunks” rather than loading it all at once.
Example: Process Large Transaction Logs
import pandas as pd
file_path = 'large_transactions.txt'
# Setup a chunk size (number of rows per iteration)
chunk_size = 1000
# This returns an iterable object
reader = pd.read_csv(file_path, chunksize=chunk_size)
for chunk in reader:
# Process each chunk (e.g., filter rows or calculate sums)
print(f"Processing chunk with {len(chunk)} rows...")
# For this example, we'll just break after the first chunk
breakThis “streaming” approach is the professional way to handle Big Data on a standard laptop.
Common Troubleshooting Tips
Over the years, I’ve run into a few recurring issues when reading text files. Here is how I solve them:
- Encoding Errors: If you see strange characters, try adding encoding=’utf-8′ or encoding=’latin1′.
- Missing Values: If your file uses “n/a” or “Empty” instead of staying blank, use na_values=[‘n/a’, ‘Empty’].
- Trailing Spaces: If your column names have accidental spaces (e.g., “Price “), use skipinitialspace=True.
In this tutorial, we looked at several ways to read text files using Pandas, from basic CSV-style formatting to handling fixed-width legacy files.
I’ve found that mastering the sep, skiprows, and usecols parameters covers almost every real-world scenario you’ll face.
You may also like to read:
- Ways to Set Column Names in Pandas
- How to Get Length of DataFrame in Pandas
- How to Display All Columns in a Pandas DataFrame
- How to Export Pandas DataFrame to CSV in Python

Bijay Kumar is an experienced Python and AI professional who enjoys helping developers learn modern technologies through practical tutorials and examples. His expertise includes Python development, Machine Learning, Artificial Intelligence, automation, and data analysis using libraries like Pandas, NumPy, TensorFlow, Matplotlib, SciPy, and Scikit-Learn. At PythonGuides.com, he shares in-depth guides designed for both beginners and experienced developers. More about us.

