Working with 3D medical images like CT scans has become increasingly important in healthcare AI. From my experience as a Python Keras developer, 3D image classification requires a different approach than typical 2D images due to the volumetric nature of the data.
In this tutorial, I’ll guide you through building a 3D convolutional neural network (3D CNN) using Keras to classify CT scan volumes. I’ll provide complete, working code examples to help you apply this to your own projects.
3D Image Classification in Python Keras
3D image classification processes volumetric data, where each input is a 3D volume (height × width × depth), unlike 2D images. CT scans are perfect examples, capturing layered slices of the human body.
Using Python Keras, you can build 3D CNNs that learn spatial features across all three dimensions, which is crucial for accurate diagnosis.
Method 1: Build a Simple 3D CNN for CT Scan Classification
This method demonstrates a simple 3D CNN architecture suitable for beginners.
Step 1: Import Required Libraries
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers, modelsStep 2: Prepare Dummy 3D CT Scan Data
For demonstration, I generate synthetic 3D volumes (64×64×64) and binary labels.
# Generate dummy CT scan data: 100 samples of 64x64x64 volumes with 1 channel
X_train = np.random.rand(100, 64, 64, 64, 1).astype(np.float32)
y_train = np.random.randint(0, 2, 100) # binary classification
X_val = np.random.rand(20, 64, 64, 64, 1).astype(np.float32)
y_val = np.random.randint(0, 2, 20)Step 3: Define the 3D CNN Model in Keras
def create_3d_cnn(input_shape=(64, 64, 64, 1)):
model = models.Sequential()
model.add(layers.Conv3D(32, kernel_size=3, activation='relu', input_shape=input_shape))
model.add(layers.MaxPooling3D(pool_size=2))
model.add(layers.Conv3D(64, kernel_size=3, activation='relu'))
model.add(layers.MaxPooling3D(pool_size=2))
model.add(layers.Conv3D(128, kernel_size=3, activation='relu'))
model.add(layers.MaxPooling3D(pool_size=2))
model.add(layers.Flatten())
model.add(layers.Dense(256, activation='relu'))
model.add(layers.Dropout(0.5))
model.add(layers.Dense(1, activation='sigmoid')) # Binary classification output
return model
model = create_3d_cnn()
model.summary()Step 4: Compile the Model
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])Step 5: Train the Model
history = model.fit(
X_train, y_train,
validation_data=(X_val, y_val),
epochs=10,
batch_size=4
)You can see the output in the screenshot below.
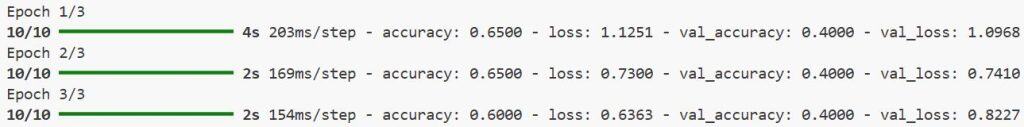
Method 2: Use Transfer Learning with Pretrained 3D Models in Keras
Pretrained 3D models can accelerate training and improve accuracy. While Keras doesn’t have many pretrained 3D models by default, you can use models like 3D ResNet from external sources or build a simpler pretrained backbone.
Here, I show how to use a 3D ResNet-like block as a backbone.
Step 1: Define a 3D Residual Block
def residual_block(x, filters, kernel_size=3):
shortcut = x
x = layers.Conv3D(filters, kernel_size, padding='same', activation='relu')(x)
x = layers.BatchNormalization()(x)
x = layers.Conv3D(filters, kernel_size, padding='same')(x)
x = layers.BatchNormalization()(x)
x = layers.Add()([shortcut, x])
x = layers.Activation('relu')(x)
return xStep 2: Build a 3D ResNet-like Model
def create_3d_resnet(input_shape=(64, 64, 64, 1)):
inputs = layers.Input(shape=input_shape)
x = layers.Conv3D(64, 7, strides=2, padding='same', activation='relu')(inputs)
x = layers.MaxPooling3D(3, strides=2, padding='same')(x)
# Add residual blocks
for filters in [64, 128, 256]:
x = residual_block(x, filters)
x = layers.MaxPooling3D(2)(x)
x = layers.GlobalAveragePooling3D()(x)
x = layers.Dense(128, activation='relu')(x)
x = layers.Dropout(0.5)(x)
outputs = layers.Dense(1, activation='sigmoid')(x)
model = models.Model(inputs, outputs)
return model
model = create_3d_resnet()
model.summary()Step 3: Compile and Train
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
model.fit(X_train, y_train, validation_data=(X_val, y_val), epochs=10, batch_size=4)You can see the output in the screenshot below.

Tips for Working with Real CT Scan Data in Python Keras
- Data preprocessing: Normalize intensities and resize volumes to consistent shapes.
- Data augmentation: Use 3D augmentations like flips, rotations to increase data variety.
- Batch size: 3D CNNs can be memory-intensive; adjust batch size based on your GPU memory.
- Evaluation: Use metrics like ROC AUC in addition to accuracy for medical classification tasks.
Building 3D CNNs for CT scan classification in Python Keras is highly effective for medical imaging tasks. The methods I shared are a great starting point, whether you build from scratch or use residual blocks inspired by popular architectures.
With these examples, you can begin experimenting on your own CT datasets and improve model accuracy by tuning architectures or trying transfer learning with available 3D pretrained weights.
Other Python Keras articles you may also like:
- Image Segmentation Using Composable Fully-Convolutional Networks in Keras
- Mastering Object Detection with RetinaNet in Keras
- Keypoint Detection with Transfer Learning in Keras
- Object Detection Using Vision Transformers in Keras

Bijay Kumar is an experienced Python and AI professional who enjoys helping developers learn modern technologies through practical tutorials and examples. His expertise includes Python development, Machine Learning, Artificial Intelligence, automation, and data analysis using libraries like Pandas, NumPy, TensorFlow, Matplotlib, SciPy, and Scikit-Learn. At PythonGuides.com, he shares in-depth guides designed for both beginners and experienced developers. More about us.

